Boston Children's Hospital · Harvard Medical School
Cohen Laboratory of Translational Neuroimaging
We use causal neuroimaging — brain lesions and pharmaco-fMRI — to map the circuits behind autism and ADHD symptoms, then pilot non-invasive neuromodulation (TMS and real-time fMRI neurofeedback) to support rapid clinical trials.
-
Map
lesions & pharmaco-fMRI
-
Validate
in non-lesional patients
-
Modulate
TMS & neurofeedback
-
Assess
change in behavior
-
Translate
rapid clinical trials
An iterative loop — bedside to clinical trials, and back
From our work
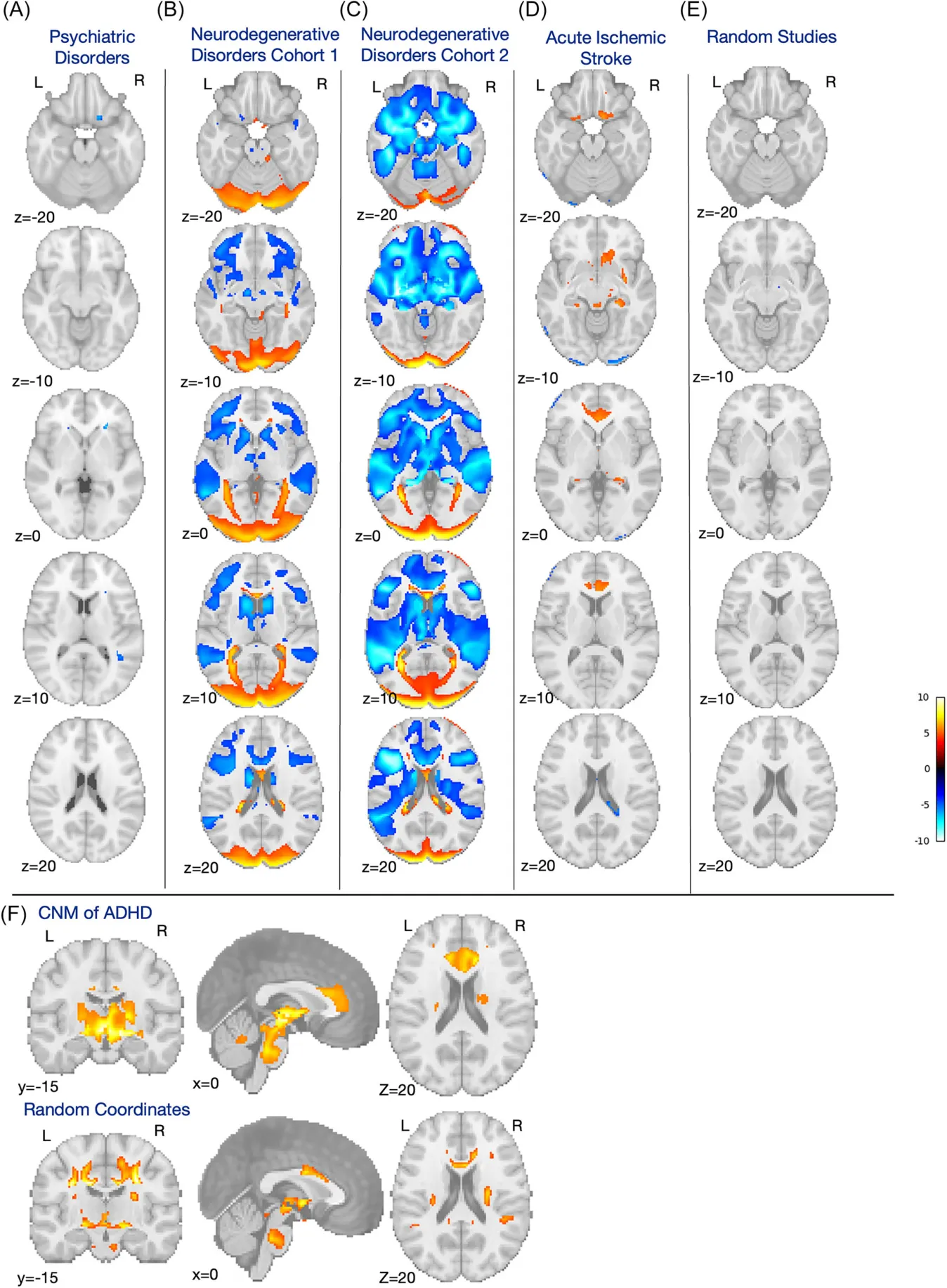
Annals of the Child Neurology Society · 2025
Coordinate Network Mapping of Focal Brain Volume Differences in ADHD Reveals Common Patterns That Lack Specificity: A Systematic Review
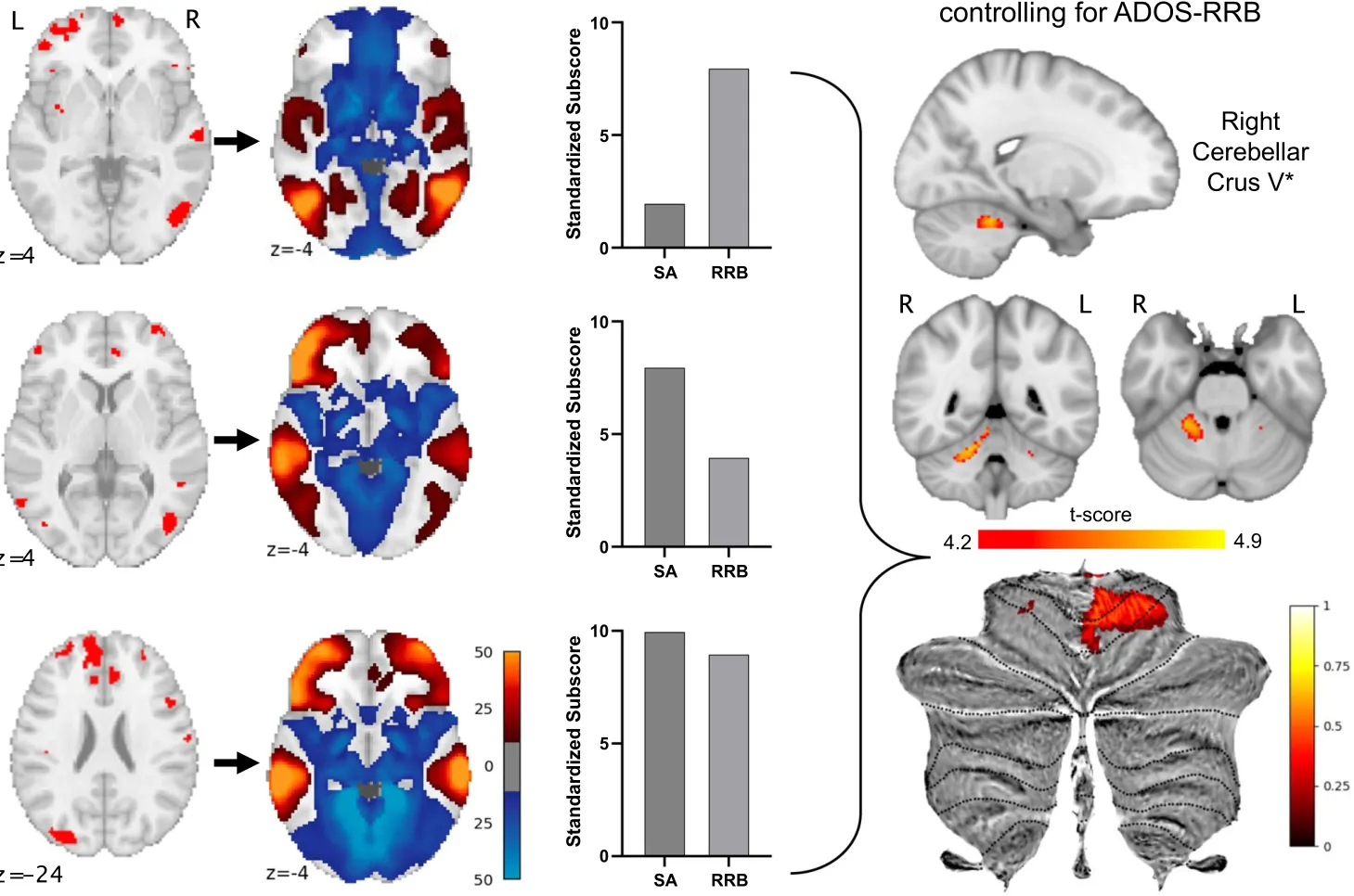
Annals of the Child Neurology Society · 2025
Lesions associated with autism symptoms map to a cerebellar brain network in tuberous sclerosis complex
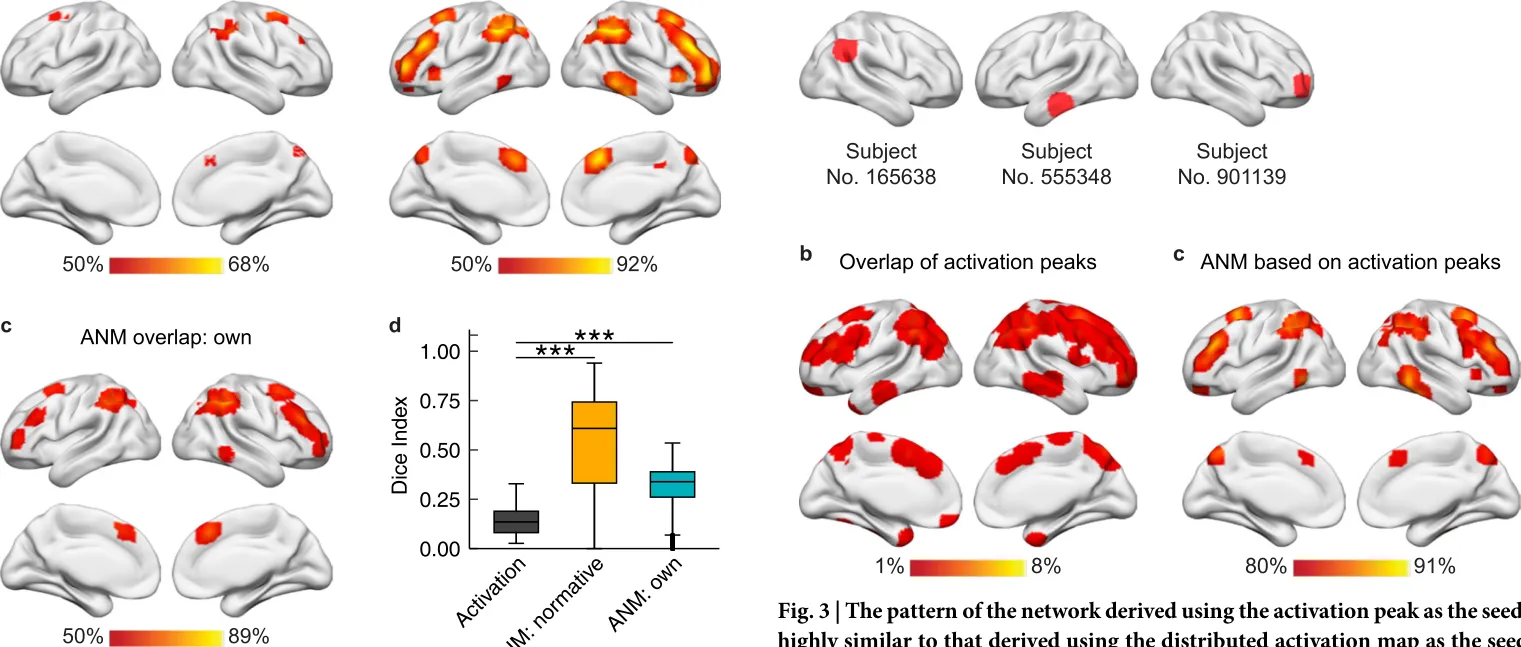
Communications Biology · 2024
Heterogenous brain activations across individuals localize to a common network
Research areas
Autism spectrum disorder
Localizing the circuits behind autism's core and associated symptoms — social communication, sensory differences, face processing — and testing targeted, non-invasive ways to modulate them.
ADHD & attention
Coordinate and lesion network mapping of attention, with pharmaco-fMRI and real-time fMRI neurofeedback to probe and modulate the circuits involved.
Tuberous sclerosis & epilepsy
Tubers, perinatal strokes, and epilepsy foci are natural experiments — focal anomalies that, when they share a symptom, reveal the responsible brain network.
Methods & open tools
Reproducible neuroimaging tools, BIDS-standard pipelines, and large-scale data harmonization — the open resources that make this work possible.
Recent work
-
Lesion network mapping of focal injury-related aggression finds two distinct network injury patterns
Brain Communications · 2026
-
Imaging Neuroscience · 2026
-
Imaging Neuroscience · 2025
-
Annals of the Child Neurology Society · 2025
-
Annals of the Child Neurology Society · 2025
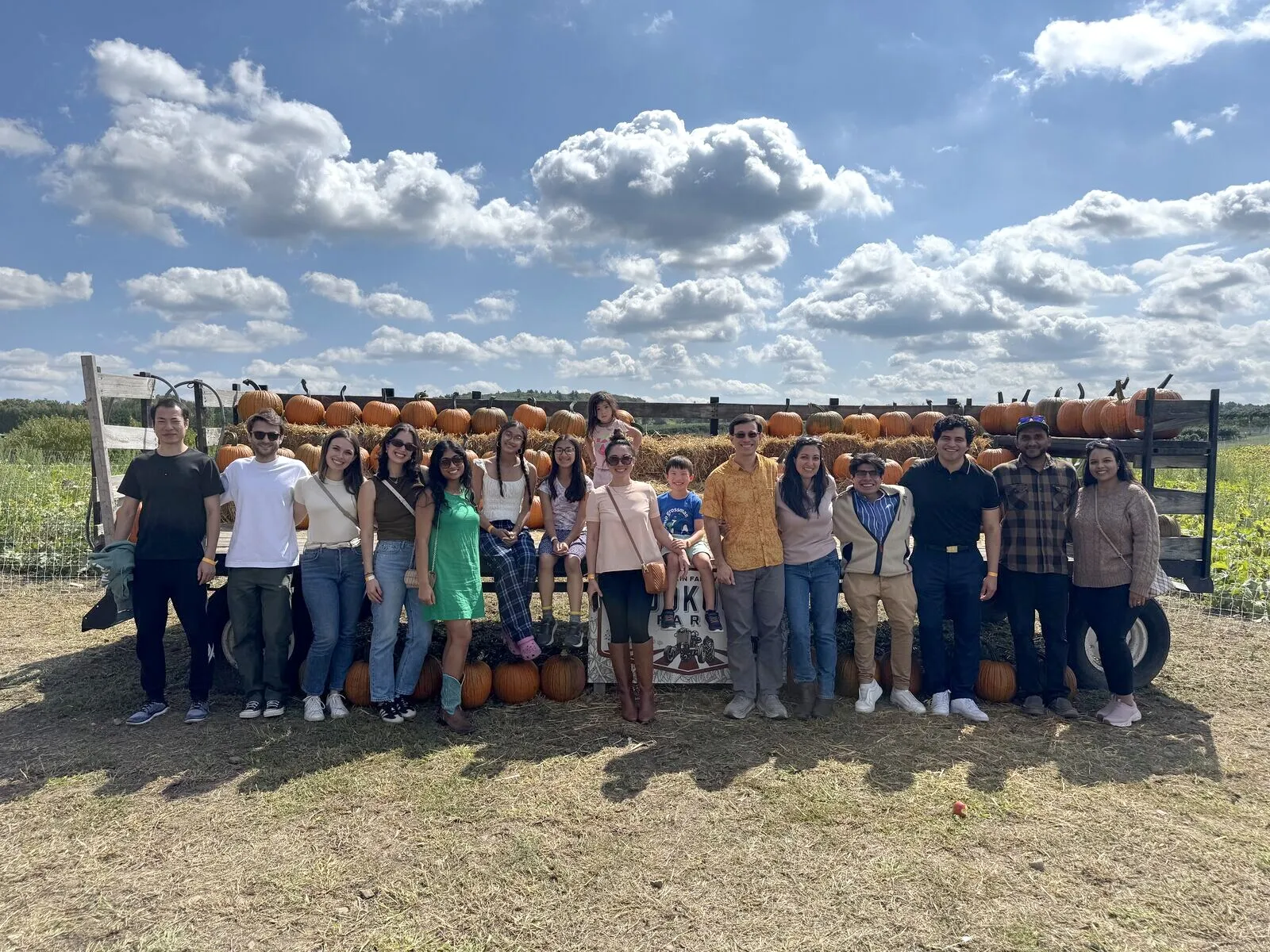
Life in the lab
Science is a team sport. Beyond the bench, we celebrate milestones, travel to conferences, and get out together — like this fall trip to the pumpkin patch.
Explore lab life











